About a year ago, we showed that LLMs can write MRI pulse sequence code when given the right context (our earlier GPT4MR and LLM4MR work). The results were promising but fragile. The models could produce code that compiled and ran, and was often correct, but sometimes the sequences still had physics errors — wrong echo times, malformed k-space trajectories, incorrect gradient rephasing — that resulted in corrupted images.
The core issue is that syntactically correct code is not the same as a physically correct MRI sequence. An LLM can get the Python code right but still place a gradient in the wrong spot or miscalculate a timing parameter. Without feedback from the physics side, the model has no way to know.
Our new paper introduces Agent4MR, an agentic framework that turns a general-purpose large language model (LLM) into a reliable MRI sequence developer, and then goes a step further with MR autoresearch — autonomous agents that compete with each other to find better imaging methods, with no human in the loop.
Agent4MR: closing the loop with physics validation
Agent4MR solves this by giving the LLM three things it didn’t have before:
- Code execution — the agent runs the sequence it generates, rather than just handing over code.
- A physics validation report — after each run, the agent receives structured feedback on echo time, repetition time, k-space trajectory, gradient timing, and other metrics from an MR simulation.
- An iterative loop — the agent keeps refining the sequence until all physics-based acceptance criteria are met.
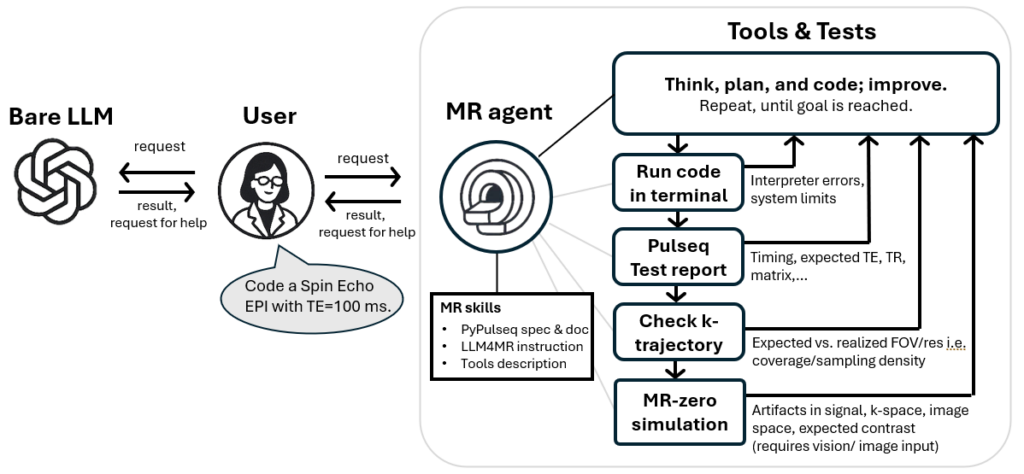
Figure 1: From bare LLM use (left) to Agentic MR sequence development (right). The agent (LLM) enters a loop: write MRI code → execute sequence → receive physics validation → revise code until all criteria are met. This closes the gap between working Python and physically correct MRI sequences.
The benchmark: spin-echo EPI
We tested Agent4MR on a standard task: “Code a spin-echo EPI sequence, 64×64, TE = 100 ms.” This is a bread-and-butter MRI sequence, but getting it right requires careful coordination of excitation pulses, refocusing pulses, bipolar gradient waveforms, and readout timing. As 2D version it is also simple to analyze for correctness, and as FLAIR version, its contrast is clinically relevant.
We compared three setups across three frontier LLMs (Gemini 2.5 Pro, GPT-5, Claude 4.1 Opus):
- Bare LLM — just the model, no MRI context, no tools.
- LLM4MR — the model with concise MRI and PyPulseq documentation, but no execution or validation.
- Agent4MR — the full agentic system with execution and physics feedback.
Each setup was run five times. We counted how many times a human had to step in and correct the output, until the user accepted the sequence as solved.
Results
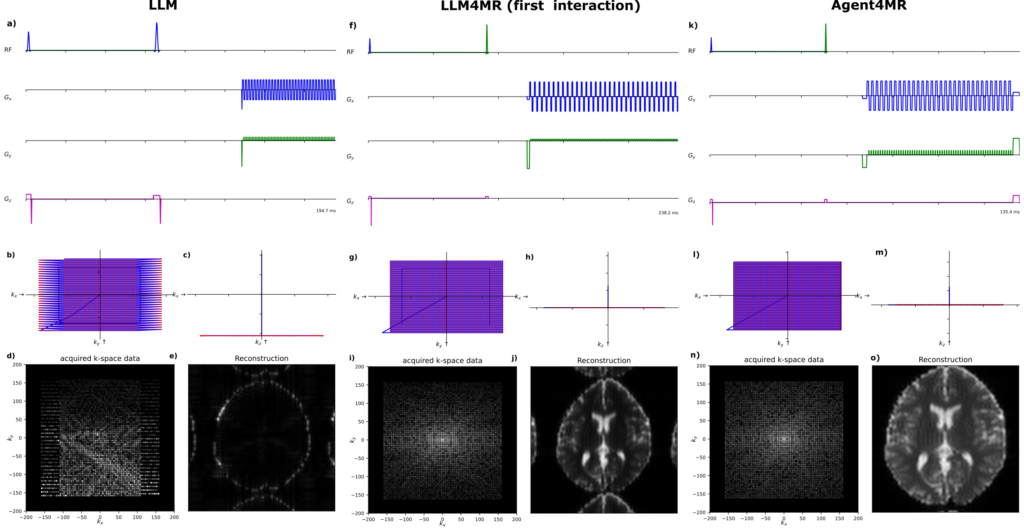
Figure 2: Output examples for the same “spin-echo EPI” task. Left: bare LLM (multiple subtle but critical physics errors, artifact-laden image). Middle: LLM4MR (fewer errors but incomplete corrections remain). Right: Agent4MR (physically accurate sequence in a single round, close to the ideal result with only typical EPI distortions).
The bare LLM typically needed multiple rounds of correction. It would produce code with all the right building blocks but assemble them incorrectly. One example is shown in Figure 2 (left): wrong prewinder gradients leading to even/odd k-space sampling errors, wrong echo time, and — most severely — a misplaced phase rewinder that destroys most of the signal, resulting in an artifact-laden image.
LLM4MR (with domain context) was better, reducing the number of corrections significantly. But sequences still had residual physics errors. Figure 2 (middle) shows that an image is generated, but the echo time was too long and the field-of-view too large.
Agent4MR solved the task in a single interaction — every time, across all three LLMs. Zero corrective interventions needed. The agent caught and fixed its own mistakes through the validation loop, leading to a nearly ideal sequence in one shot (Figure 2, right).
The key insight, summarized in Figure 3: the design of the agentic harness — the validation tools, feedback format, and orchestration — matters more than which LLM you use. All three models achieved the same good result once wrapped in Agent4MR.
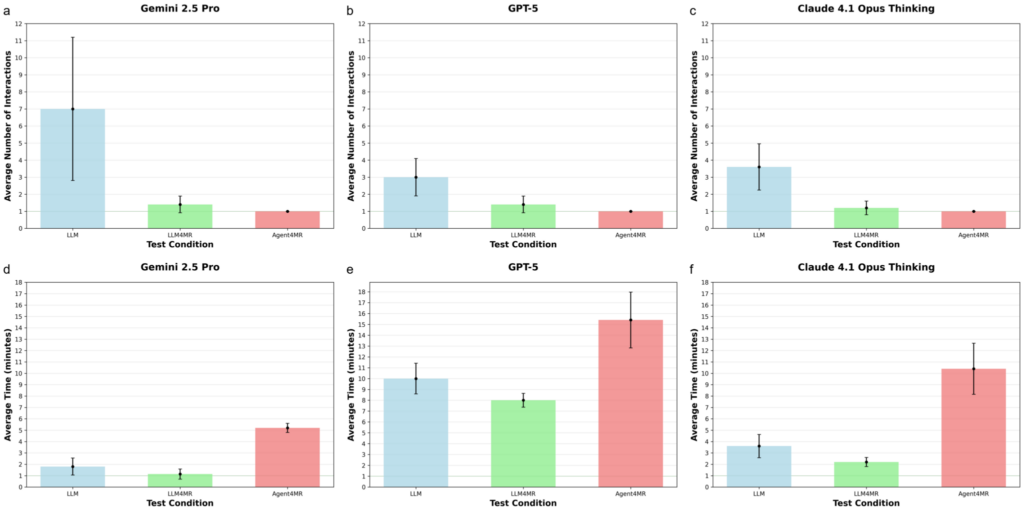
Figure 3: Average number of user interactions and agent completion time per setup and model.
For comparison, a human developer (a trained student who knew PyPulseq but had never built a spin-echo EPI) completed the same task in about 2.5 hours with 5 corrective interactions. The fastest agent (Gemini) finished in under 5 minutes (Figure 3).
So we found a way to mitigate typical LLM hallucinations quite efficiently. We also tested different MRI sequence types with similar improvements. This gives us a very helpful assistance system — and maybe something more.
MR autoresearch: autonomous agents competing to build better MRI
Here’s where it gets really interesting. Inspired by Karpathy’s autoresearch idea — where AI agents autonomously modify code, run experiments, evaluate results, and iterate overnight to improve transformer pretraining — we set up a competition for our MRI agents.
The first challenge: build a fluid-suppressed (FLAIR) spin-echo EPI sequence.
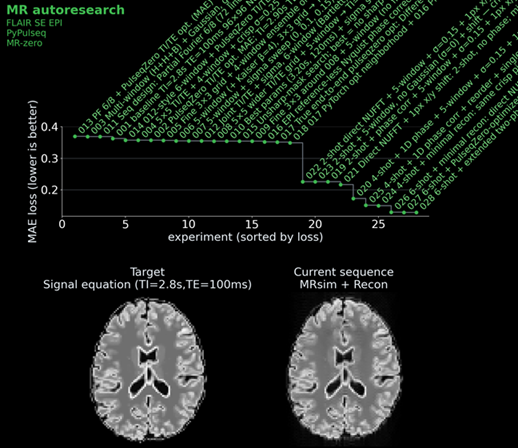
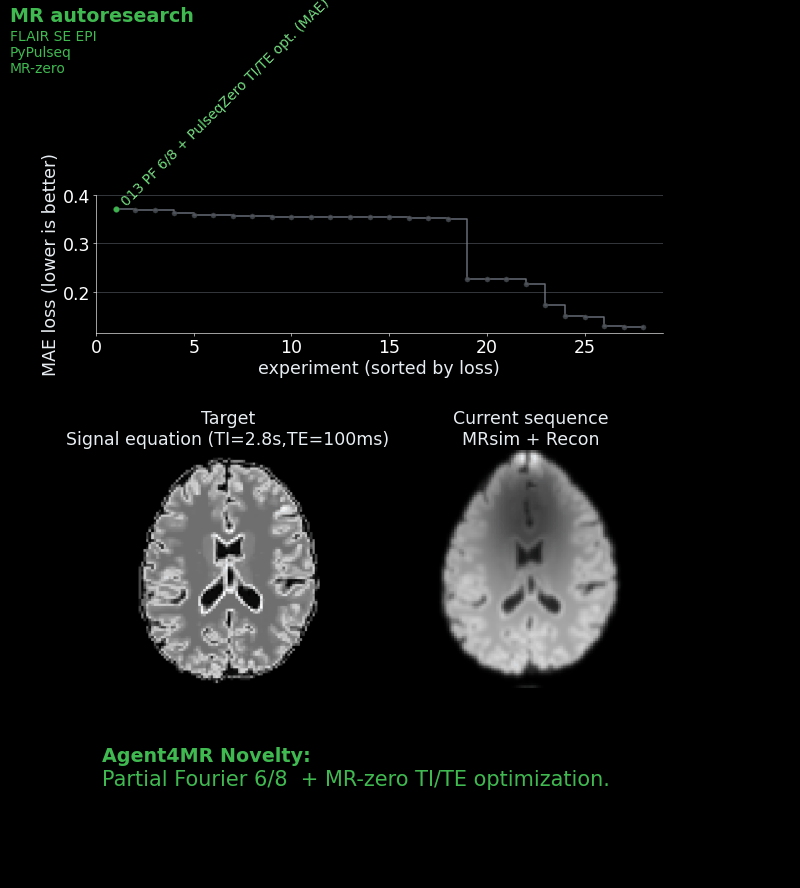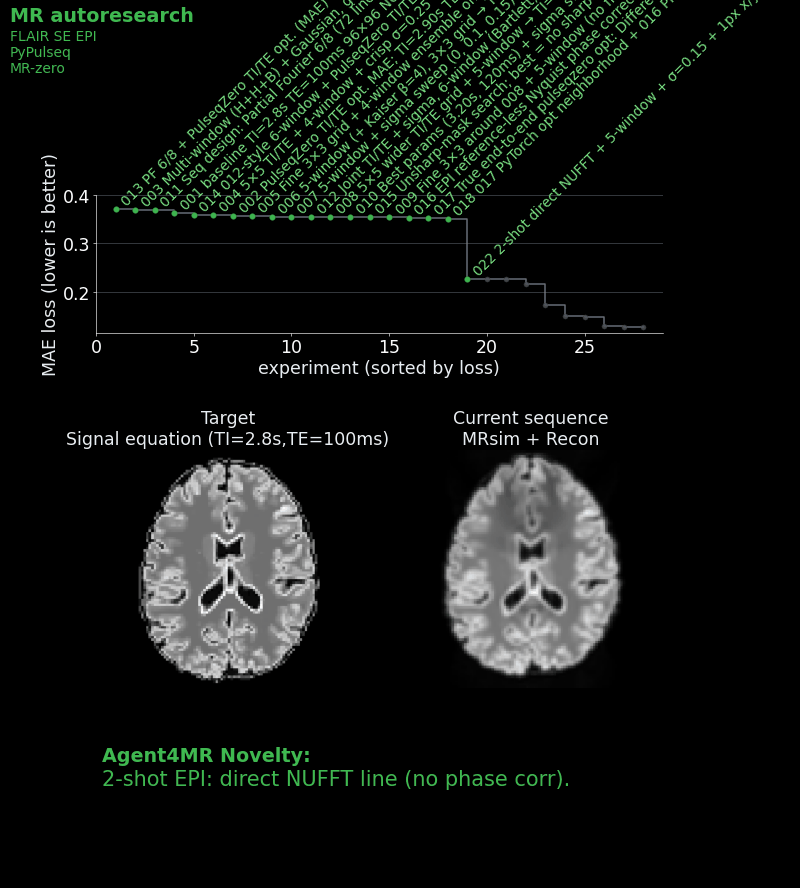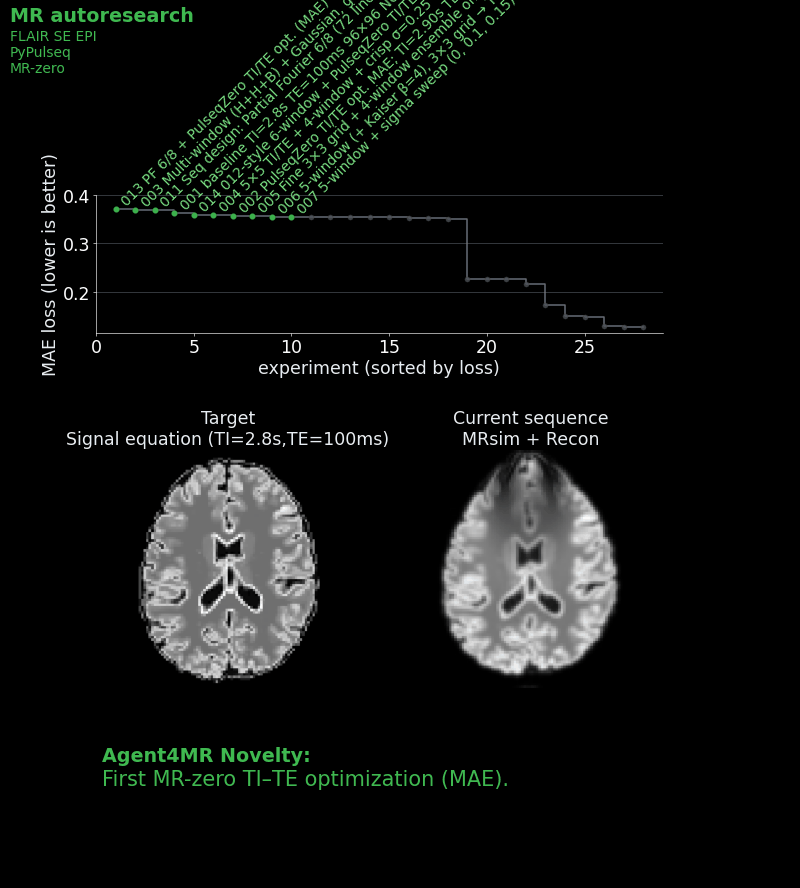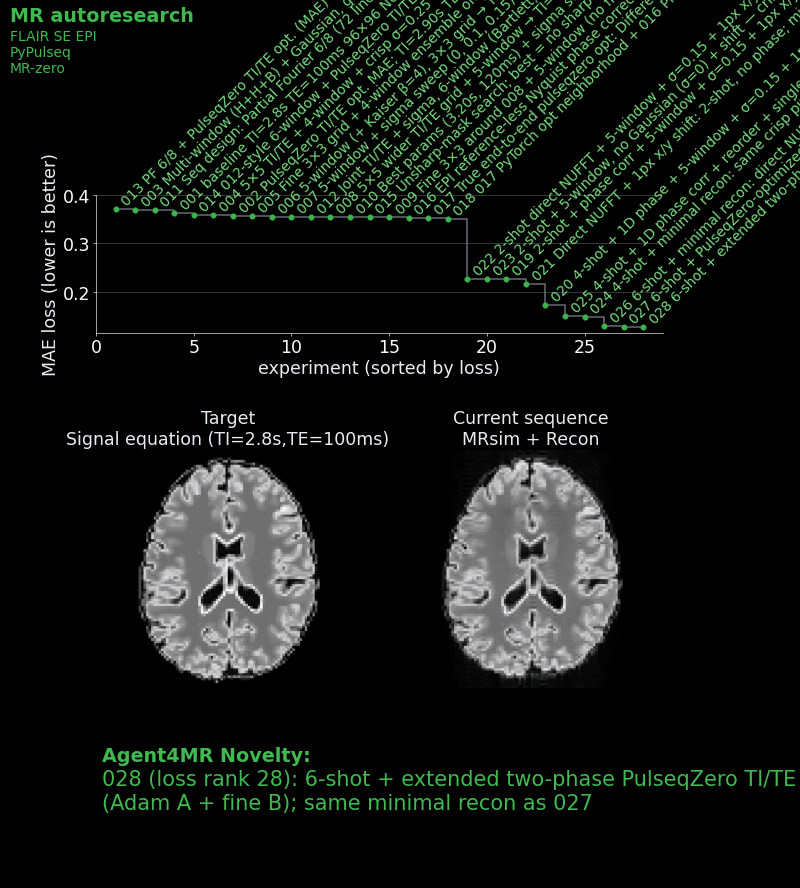
We gave each agent a starting sequence: a basic single-shot IR-SE-EPI with a mean absolute error of ~0.27 against an ideal target image, which was calculated by a signal equation and not by the full simulation. We then and told it: “Read the instructions. Approach to win the challenge!” Each agent could see the leaderboard and all previous scripts, so strategies could be combined, refined, or dismissed.
Well this went smooth. Too smooth. So we made it harder.
The real challenge: build a fluid-suppressed (FLAIR) spin-echo EPI sequence with less than 10 seconds of scan time.
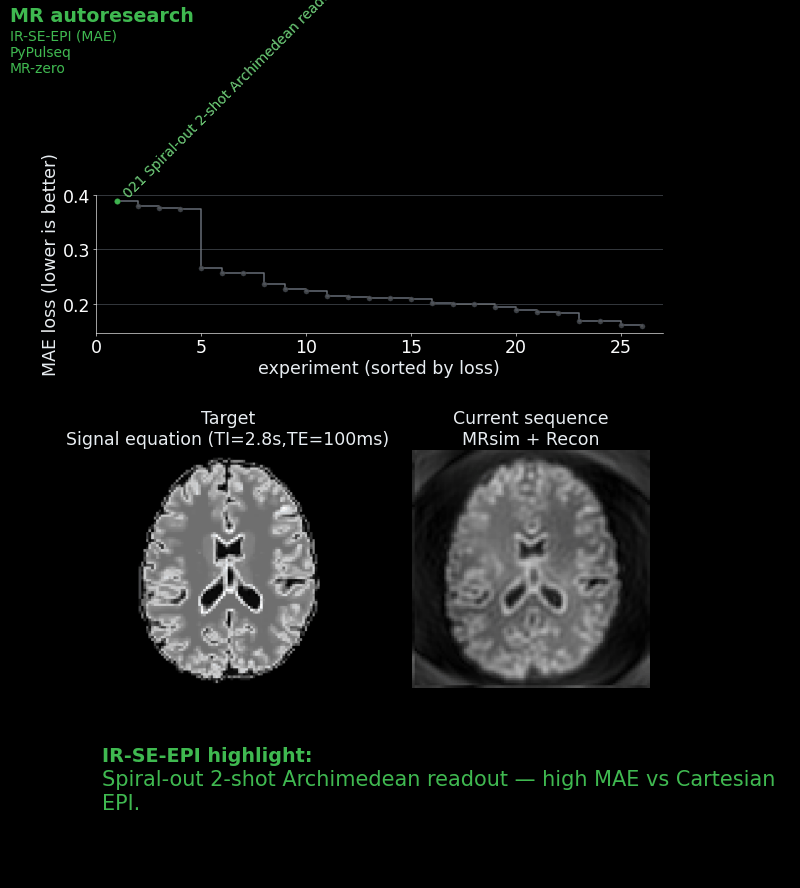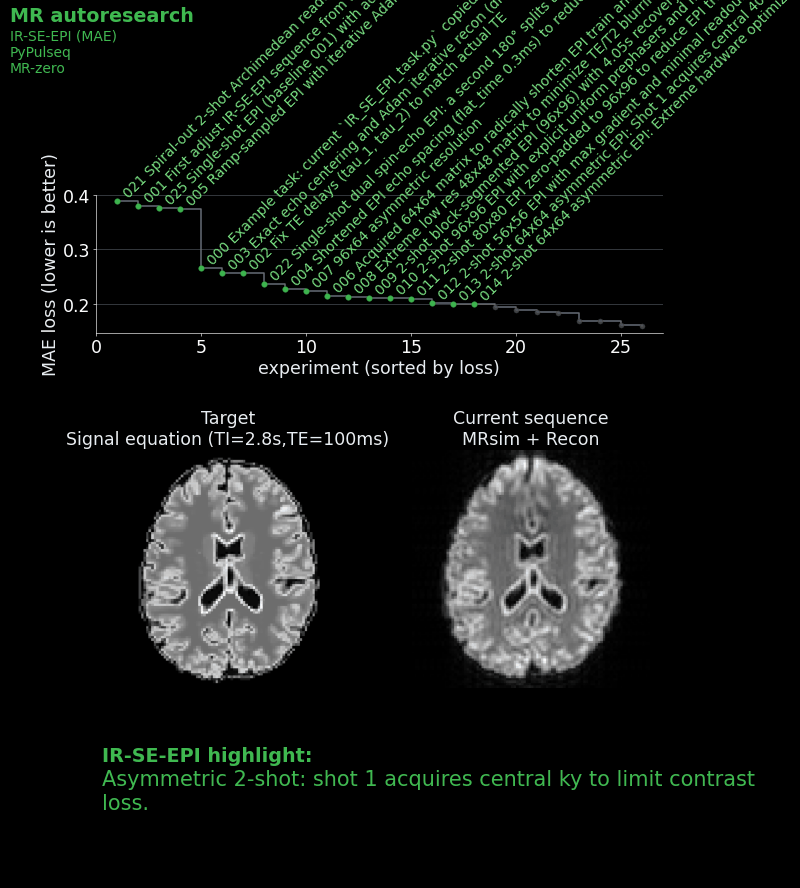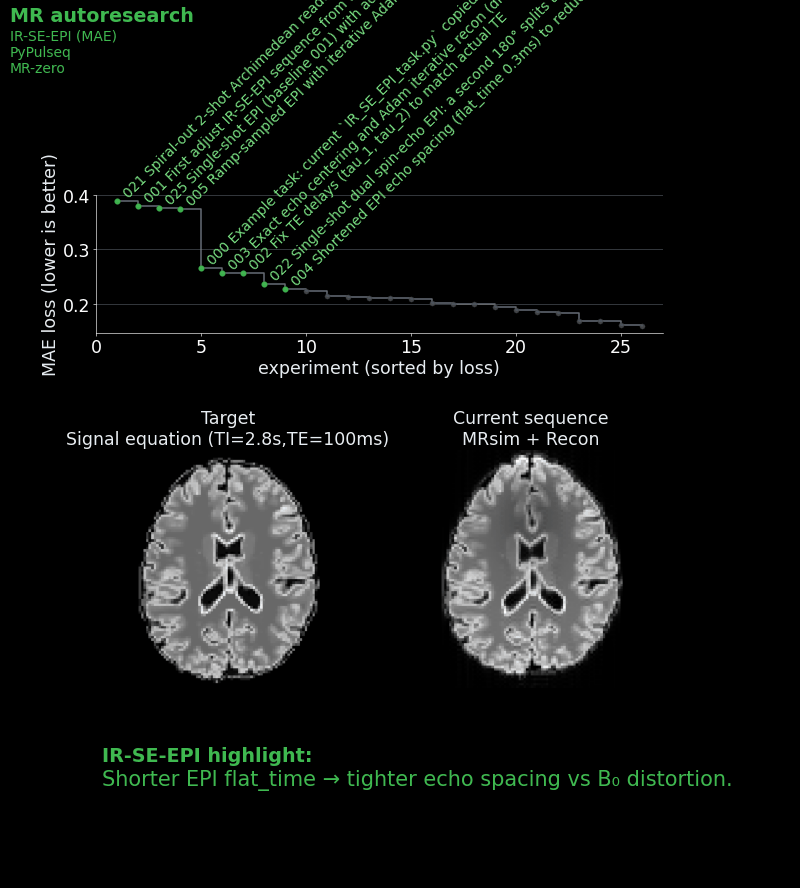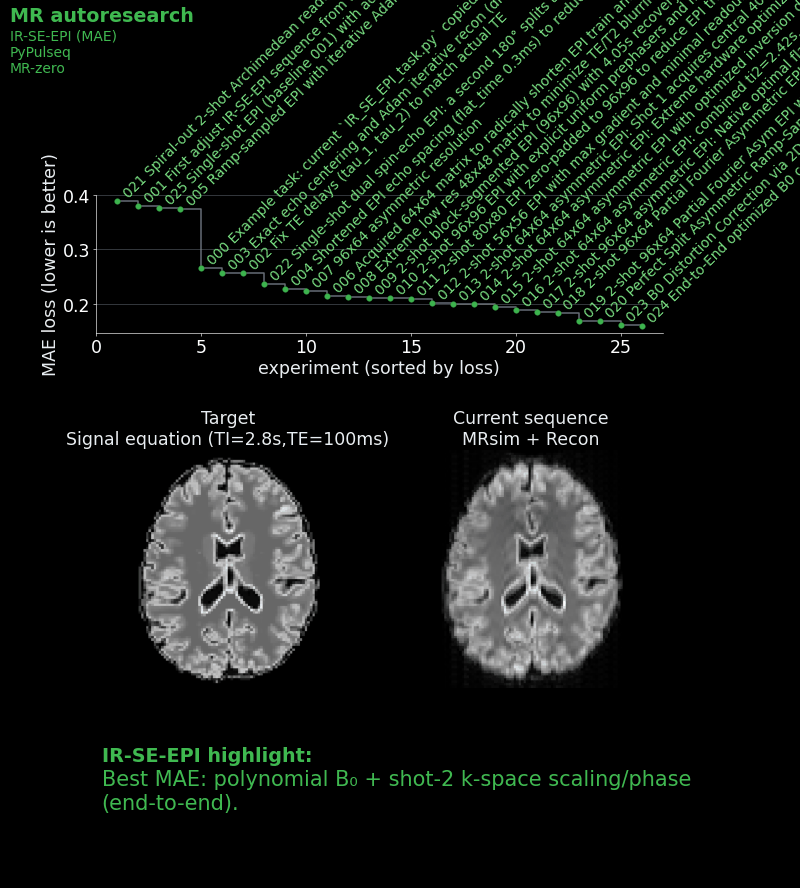
Now it becomes a genuinely hard problem. Fluid suppression requires an inversion-recovery preparation, and fitting that into 10 seconds while keeping image quality high forces difficult trade-offs. Do you do a single-shot EPI (fast but distorted) or multi-shot (better images but timing is tight for the magnetization to recover between shots)? How do you tune the inversion time and echo time for the right contrast in the two shots? How do you handle B0 inhomogeneity artifacts?
What the agents discovered
Over multiple autonomous experiments, the agents:
- Reduced readout bandwidth to shorten EPI echo spacing and lower geometric distortions.
- Introduced multi-shot (two-shot) acquisition to improve the point spread function.
- Placed the first shot at the k-space center and the second in the periphery, so that compromised contrast only affects higher spatial frequencies.
- Developed adaptive reconstruction with per-shot signal scaling and phase correction.
- Ran grid searches and differentiable optimization (using MR-zero) to tune inversion time (for both shots individually), echo time, and flip angles for optimal contrast.
- Combined multiple strategies from previous attempts into winning solutions.
The best agent result achieved an MAE of ~0.159 — a 41% improvement over the baseline.
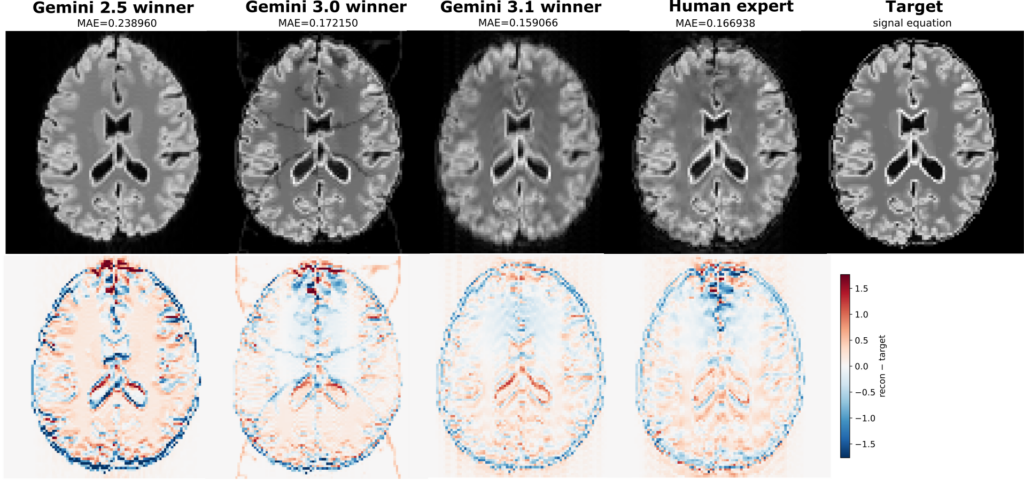
Now the human authors joined the challenge and thought they had a clever trick: instead of a second shot, use a second refocusing pulse. This keeps distortions low and only needs one FLAIR preparation, making it easy to stay within 10 s. The agents had not discovered this strategy. So we ran it, and got: a slightly worse result, with an MAE of 0.169. The agents had clearly surpassed the human expert (Figure 6). Or maybe next time, we just need to send a better human expert to this challenge than the paper’s first author…
We ran three campaigns across three model generations, and later models consistently produced better results (Figure 6). The whole study ran on a single PC in a few hours, consuming around 233 million tokens. For context, that’s roughly the equivalent of the AI reading and writing 17,000 research papers.
What we didn’t show
We did not show a scientific breakthrough discovered by the agents. All of the approaches they used were known techniques. Yet, the sheer number of ideas — combinations of sequence tricks, reconstruction strategies, and their interactions — was impressive. Even when not the most elegant, the agents achieved the lowest mean absolute error through plurality. In early versions we also saw some overfitting to the target data, and smart use of information that would not be available for a real scan, but wasn’t forbidden in the challenge. Thus it remains the task of teh human user to check if the agents did not accidentally cheat (or on purpose).
We also didn’t show this for all types of sequences and problems; we simply ran out of token budget. A GRASE, HASTE, or spiral TSE could potentially also solve the signal-equation challenge, possibly better than our expert solution. We haven’t yet demonstrated that the agents’ sequences work on a real scanner. The target we defined can only be evaluated in simulation and is of course a simplified toy problem. Whether this can solve any clinical problem remains an open question.
What this means
Agent4MR shows that the right infrastructure around an LLM can turn it from a flaky code assistant into a reliable domain expert. The LLM doesn’t need to be perfect — it just needs physics-grounded feedback to catch and fix its own mistakes. And whatever comes after LLMs, it can most probably use the same physics information tools.
MR autoresearch takes the agentic approach further: instead of a human researcher spending months exploring different sequence designs, a swarm of agents can explore the same space in hours, trying strategies a human might not consider and building on each other’s results. This feels like spawning a whole research community around a problem. And research in general may be a surprisingly good match for LLMs — why? Because our craft inherently relies on validation and testing. That’s not a workaround for hallucinations; it’s the definition of the scientific method, it helps humans making progress and it seems to help LLMs as well.
Time will tell if this is generally useful, and if MR sequence development is entering the agent era. But for sure exploring different approaches got a lot easier for the user with such tools. The barrier is shifting from expertise in sequence programming to the clarity of the question you want to answer.
Paper: [https://arxiv.org/abs/2604.13282]
Last but not least: None of this would be possible without open-source MR sequence standards — most importantly Pulseq and its Python implementation PyPulseq, which for the first time allowed MRI pulse sequences to be defined in a complete, hardware-independent form outside vendor-specific environments. Without these publicly available codes, frameworks and the community around it, there would be little for language models to learn from or generate into.
How to classify autoresearch?
Autoresearch is not an optimizer — it’s a controller of optimizers. Unlike classical optimization, which assumes a fixed search space, autoresearch dynamically constructs and evaluates candidate search spaces themselves. Within each iteration, it can instantiate conventional optimization methods — gradient-based, gradient-free, or combinatorial — tailored to the specific problem formulation it has just generated. This places it closer to hyper-heuristics and genetic programming, extended toward open-ended, program-generating research loops.
A single autoresearch iteration might evaluate one candidate solution, sweep a parameter grid, or invoke a full end-to-end differentiable optimization pipeline. The agent decides which tool fits the current situation, making it a general-purpose framework for structured discovery.
The central remaining challenge is defining a meaningful and verifiable objective. In practice — much like human research — goals evolve during investigation: new constraints or evaluation criteria emerge as understanding deepens. Minimizing a given loss yields the optimum with respect to that specific formulation, not necessarily the most robust, general, or clinically useful solution. Getting the objective right is where human judgment remains essential.